AGS/ALS/cGAS to Parkin/PRKN Transition
AGS/cGAS Evidence Table Continuation
The visible top rows are the lower part of a larger AGS/cGAS table. The partial column/header text includes:
accumulation of cytosolic self-DNA (exonuclease TREX1)
c self-DNA (mito DNA아님)
(cGAMP) 생성함
STING
IRF3
response genes in the nucleus
T-Cell Activation| block | model / evidence | visible note |
|---|---|---|
Evidence / Human | no visible entry in this crop | Empty row or content above the crop. |
Evidence / Mice | {Gao, 2015 #772} TREX1-deficient mice | In heart tissue, kidney and PBMC, sera: cytosolic accumulation of self-DNA (이건 논문에 없는 느낌?) and cGAMP (in heart tissue, using MS); ↑ mRNAs of ISGs (IFN-stimulated genes, by RT-PCR (qRT-PCR)), including IFIT3, CXCL10 (=IP-10), and IRF7, as well as the inflammatory cytokines TNF-a and IL-12. |
Correction / mice | {Gao, 2015 #772} genetic ablation of cGas in Trex1-/- mice | Eliminated all detectable pathological and molecular phenotypes, including ISG induction, autoantibody production, aberrant T-cell activation, and lethality. Even deletion of just one allele of cGas largely rescued the Trex1-/- phenotypes of mice. |
primary culture | {Vincent, 2017 #773} bone marrow-derived macrophages (BMDMs) from Trex1-/- mice | RU.521 reduces IFNb1, IL-6 mRNA, in the presence of immune stimulators: dsDNA, 5'ppp-HP20 RNA, Pam3CSK4, poly(I:C), LPS, or recombinant murine interferon-beta. |
In vitro | {Vincent, 2017 #773} RAW cells | RU.521 reduces IFNB-1 (type I IFN) in RAW cells. |
ALS with cGAS
Rationale
(Neta's email 20200201)elevated levels of the specific cGAS signalling metabolitecGAMPin iPSC-derived motor neurons and post-mortem spinal cord samples of patients with ALS.
Mechanism
(Neta's email 20200201)accumulation of wild-typeTDP-43and further exacerbated by mutantTDP-43.- Mislocalized
TDP-43invades mitochondria viaTIM22and causes the mitochondrial Permeability Transition Pore (mPTP) to release mitochondrial DNA (mtDNA) into the cytoplasm. mtDNAdirectly activatescGAS→↑ cGAMP→↑ STING→ upregulation ofNF-kBand type I IFN → neuron loss.
Progress
MNIis testing(neuronal cell death assay)our compound in 3D coculture of motor neuron(ALS patient-derived)with astrocyte.Yan; Neuronal cell death assay with ALS iPSC NM in EONU which is the same with MAP4K4/ferroptosis dual inhibitor programWill get results by Mar 2021
HD with cGAS
Will be challenging
Parkin / PARK2 Gene (=PRKN)
Location
- Chromosome 6 in human
- Chromosome 17 in mouse
- Codes for the ubiquitin E3 ligase that is involved in …
PRKNgene is located at6q26, consists of12 exons
Position
- Cytogenetic location:
6q26 - Genomic coordinates (
GRCh38):6:161,347,416-162,727,801(from NCBI) - Human
PRKNhas8 mRNA transcript variants - Variant 1, a major transcript,
CDS used in program,1398 bpin size, and encodes Parkin protein isoform 1
Expression
- Parkin is ubiquitously expressed
- Parkin is highly expressed throughout the brain including
SN:https://www.genecards.org/cgi-bin/carddisp.pl?gene=PRKN - Parkin is the most abundant in neurons in the human brain
(Jacobo, Sarah Melissa) - Expressed in heart, testis and skeletal muscle
- Found in serum:
https://www.genecards.org/cgi-bin/carddisp.pl?gene=PRKN
PRKN Gene, Homology, and Expression Figure
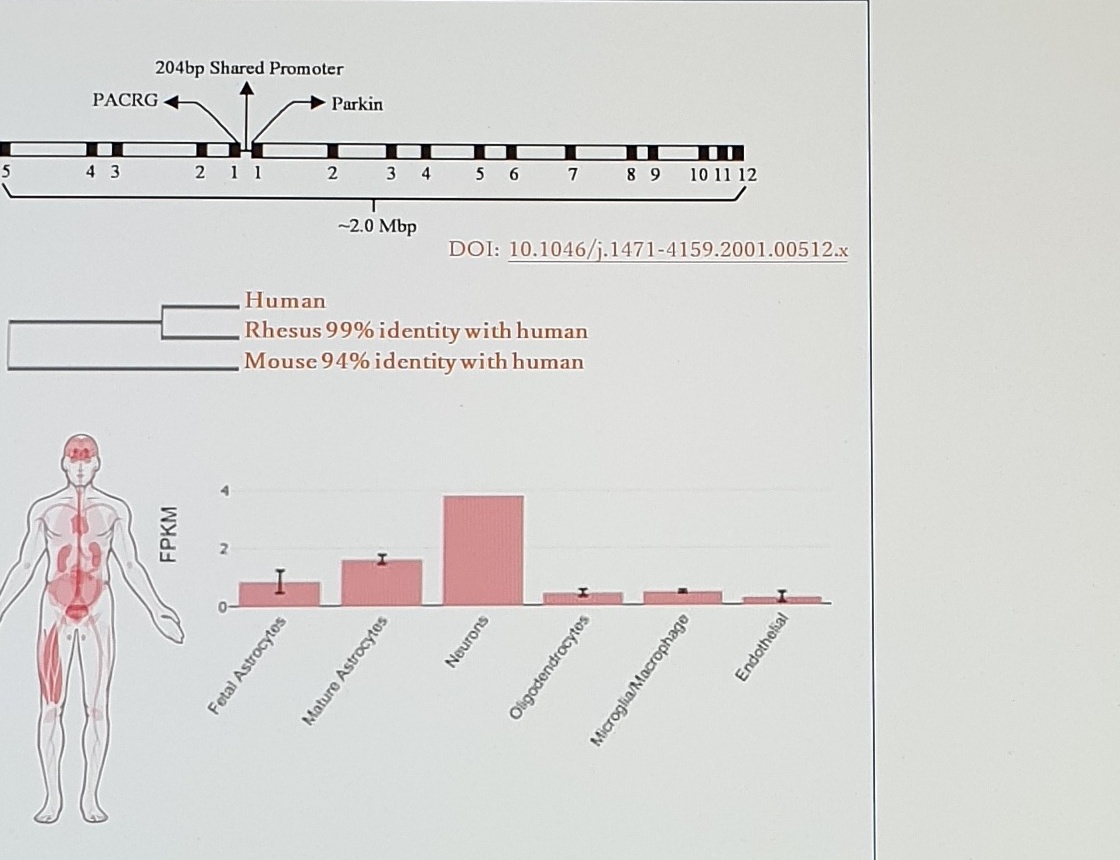
204bp Shared Promoter
PACRG
Parkin
~2.0 Mbp
DOI: 10.1046/j.1471-4159.2001.00512.x
Human
Rhesus 99% identity with human
Mouse 94% identity with human
FPKM
Fetal Astrocytes
Mature Astrocytes
Neurons
Oligodendrocytes
Microglia/Macrophage
EndothelialSplicing
- Human
PRKNhas26 splice variants:https://www.ncbi.nlm.nih.gov/pmc/articles/PMC4124806/ - Variant 1 is the major transcript with
1398 bp CDS - Variant 1 coding sequence (
CDS) is used in this program - Variant 1 encodes for Parkin protein isoform 1
Parkin Protein / Concentration of Parkin Protein
The table is partially cut at the left and bottom. Visible column groups:
Human: Brain / CSF / serum
NHP: Brain / CSF / serum| source / method | Human brain | Human CSF | Human serum | NHP brain | NHP CSF | NHP serum |
|---|---|---|---|---|---|---|
Takeshi from the commercially available human brain / ELISA | 771 (500-1500) pg/mg total protein | blank visible | blank visible | blank visible | blank visible | blank visible |
TM: SMCxPro-/well; capture antibody: antiParkin abcam, rabbit mono; EPR18567 176 12.5 ug/MP mg, 5 ugMP/well; detection antibody: antiParkin abcam, rabbit mono; EPR18567 214 125 ng/mL | SMCxpro; (n=5) 1.3 to 22.7 pg/mg total protein; Mean: 6.92 pg/mg (=7ng/g=0.007ug/g) 적네!; SD: 8.894; Warning: sample size is less than 30, you should be using the POPULATION standard deviation.; 80% CI: 6.92 +/- 5.1 (1.82 to 12); "With 80% confidence the population mean is between 1.82 and 12, based on only 5 samples."; Cf) {Quinn, 2012 #2128} aSyn (LC-MS): 500 ug/g protein | SMCxpro; n 7; Minimum 0.7000; Maximum 2.500; Range 1.800; Mean 1.257; SD 0.5740; SEM 0.2170 | blank visible | Monkey brain sample will be evaluated using current parkin protein assay method in August to confirm whether our assay method detects monkey parkin protein or not. | SMCxpro | No signal |
MS by DMPK / LC/MS | 3590 - 9052 pg/mg protein | bottom continues below visible crop | bottom continues below visible crop | bottom continues below visible crop | bottom continues below visible crop | bottom continues below visible crop |
Uncertain Spans
| location | text/status | reason |
|---|---|---|
| AGS/cGAS table top | partial mechanism columns | The table begins above the visible crop; only lower continuation can be reconstructed here. |
| Parkin location | codes for the ubiquitin E3 ligase that is involved in ... | Sentence is cut off after involved in; no completion inferred. |
| antibody names | EPR18567 176, 12.5 ug/MP mg, EPR18567 214, 5 ugMP/well | Small assay-method text is high-risk; spacing and unit formatting were updated from the raw photo but should be checked before assay reuse. |