Sample Size Estimation And NHP Studies Current Plan
Cohort / ClinGen Continuation
The top of this photo continues content from 20240722_182033. Stable fragments:
20230327 PD GENE_Takeda:
PRKN Homozygotes/compound heterozygotes: n=45,
PRKN heterozygous: n=79Europe
Christine Klein
Less active than PDGene for patient recruitmentEUROPE
ROPAD
NCT03866603
Europe & Israel, funded by Denali, Centogene.
focus is LRRK2 but everyone is genotyped to PRKN as well (gabi, 20220927) >200.Belgium.
identified 5 subjects with PRKN mutations.
She is willing to talk to us to explore the possibility of running study.
I'll talk to Jamie tonight to see if we can integrate this with MJFF.MJF(에 PRKN 추가하라고 부탁해라)
6 (?) by Jaya
(gabi, 20220927): Another person for finding patients could be Arndt Rolfs (Arcensus) - he has a long history with Gaucher.We started curating Parkinson's genes through ClinGen, which is an NIH supported platform that is recognized by the FDA.
a major effort needs to be invested in calling PRKN mutations versus VUS.
However, LRRK2 and GBA were prioritized.
It would be extremely important if a group of scientists would curate the variants to determine which is a VUS and which is a mutation.
It may significantly change the number of carriers.
Please take a look at the panel we assembled:
https://clinicalgenome.org/affiliation/40079/.
There are researchers (Ed Fon) who are on the panel exactly for PRKN.Linked URL and document names:
http://www.pdgene.org/view?poly=rs3734464
I started to put together some questions for our Monday meeting with Roy Alcalay:
Dr. Roy Alcalay discussion 2-20-23.docx
conversation with Dr Alcalay 8-12-2021.docx
Questions for Roy Alcalay 1-4-2022 meeting.docx
Meeting with Roy Alcalay June 28 2022.docxCentogene / ROPAD fragment:
CENTOGENE's Biodatabank contains N~140 genetically confirmed PRKN-PD patients that were enrolled into the ROPAD study.Sample Size Estimation
Heading and question text:
Sample size estimation
- What does the power or magnitude of effect need to be in order to see an effect in a sample size of 15-25 people?
- Using VMAT2 (a sensitive imaging biomarker), the change of VMAT2 signal over 2 YEARS is as follows:
Mean: -19.6%, SD 9.5%, % COV: 48% (=9.5/19.6)
- What sample size is needed to see a change over 1 year? Over 2 years?
To check the below sample size is per arm,? repeated-measures t-test90% Power / One-Sided p-value < 0.05
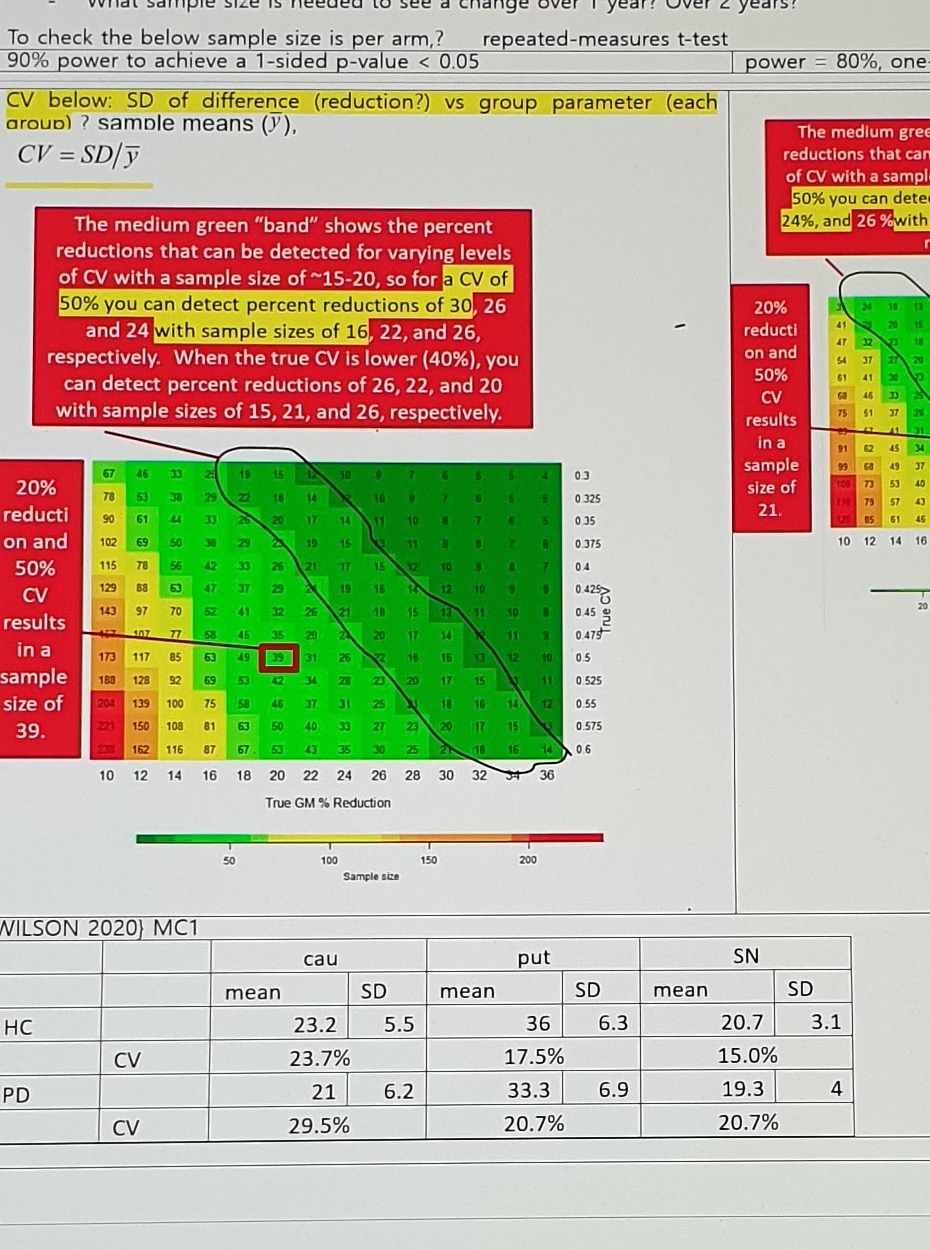
Assumption text:
90% power to achieve a 1-sided p-value < 0.05
CV below: SD of difference (reduction?) vs group parameter (each group?) sample means (y),
CV = SD / yRed-box annotation:
The medium green "band" shows the percent reductions that can be detected for varying levels
of CV with a sample size of ~15-20, so for a CV of 50% you can detect percent reductions of 30, 26
and 24 with sample sizes of 16, 22, and 26, respectively.
When the true CV is lower (40%), you can detect percent reductions of 26, 22, and 20
with sample sizes of 15, 21, and 26, respectively.Left red-box annotation:
20% reduction and 50% CV results in a sample size of 39.80% Power / One-Sided Alpha = 0.10
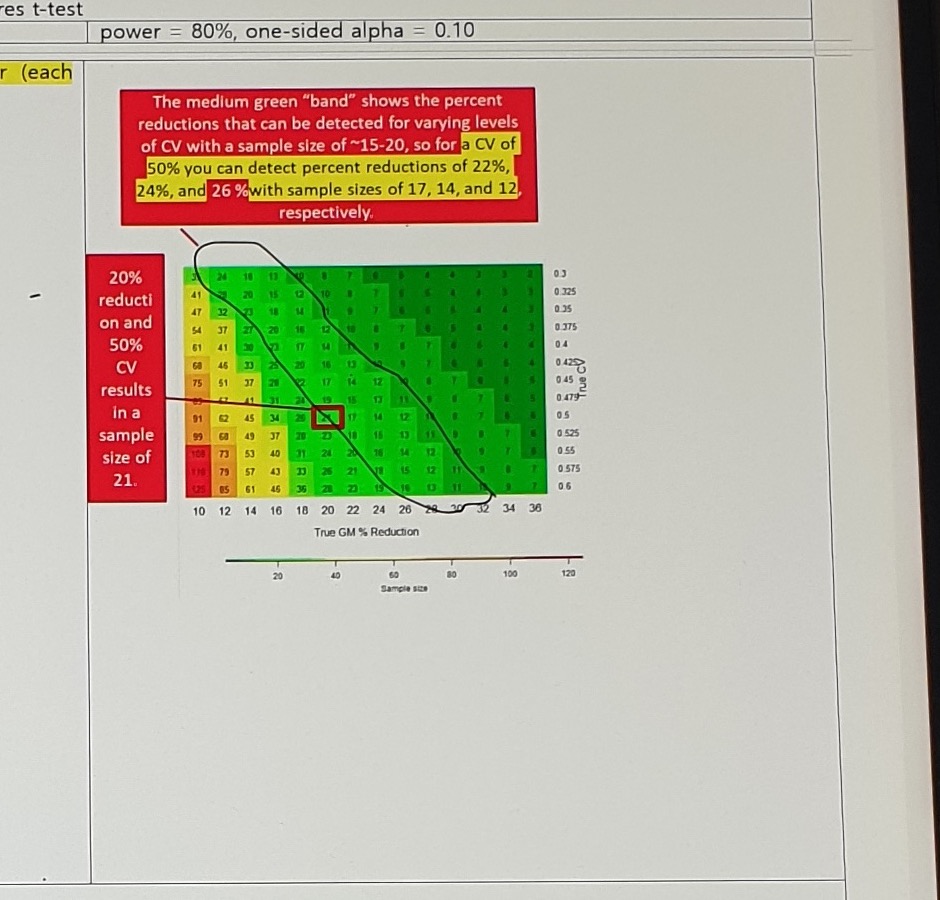
Assumption:
power = 80%, one-sided alpha = 0.10Red-box annotation:
The medium green "band" shows the percent reductions that can be detected for varying levels
of CV with a sample size of ~15-20, so for a CV of 50% you can detect percent reductions of 22%,
24%, and 26% with sample sizes of 17, 14, and 12, respectively.Left red-box annotation:
20% reduction and 50% CV results in a sample size of 21.Wilson 2020 MC1 Statistics
| Group | Statistic | cau mean | cau SD | put mean | put SD | SN mean | SN SD |
|---|---|---|---|---|---|---|---|
| HC | value | 23.2 | 5.5 | 36 | 6.3 | 20.7 | 3.1 |
| HC | CV | 23.7% | 17.5% | 15.0% | |||
| PD | value | 21 | 6.2 | 33.3 | 6.9 | 19.3 | 4 |
| PD | CV | 29.5% | 20.7% | 20.7% |
NHP Studies: Current Plan
Study Purpose / Timeline
| Study | Stable text |
|---|---|
| ROA study | ROA study = neurosurgery (a pilot study to determine the feasibility of intraparenchyal (ipa) delivery into snpc and putamen in cynomolgus monkeys |
| 1M NHP biodistribution study | targeted a Feb'23 start of study -> 4mths to have data -> CN, expecting to have it around June '23. NBR held a slot for pre-surgery MRI scans (~14 NHP) in Jan '23 and a surgery slot in Feb '23. NBR reference appears to be 047-052. |
| Toxin induced NHP study | after CN; 2023/02-2023/12; before CS; (1-3 mon in-life, 3 mon analysis); POC. |
Date milestones for the 1M NHP biodistribution study:
Injection: Feb/Mar, 2023
Necropsy: Mar/Apr, 2023
Report (IHC/ISH): Jun, 2023Promoter / Cassette Notes
ubiquitous promoter
Hattori 말들으면 astrocyte 도 중요하니까 ubiquitous promoter 사용가능성
AAV9 PK044; PGK1 as priority
*
AVV9 PK041; Syn1Dose: just highest dose possible?AVV9 PK041 may mean AAV9 PK041; the source-looking string is retained.
ROA / Dose / Criteria Notes
20221004 takeshi: @Fergus: Is there a possibility to select one ROA from two (SN and SN/putamen) by the pilot study?
This may suppress the cost of the BD study,
Based on the results of the ICM-NHP study by the GBA-PD GT project, ICM isn't in our scope now
AAV capsid / ROA evaluation
PK/PD (?) assessment and immunogenicity assessment (?)
Infusion coverage: >30%? Of SN with acceptable AE
Transgene expression: >30% neurons by IHC/ISH in SN (근데 HA-tagging 안 한다는 거지?)
No safety concernsPilot / Device Notes
1. Buffer in SNpc, N=2
2. Buffer in SNpc+Putamen, N=3
3. Buffer in SNpc+Putamen, N=3 ("Clear Point").
Cf) Current our default is a "Neurochase" device.CP)NHP Route / Dose Table
| Group | Route | Number of Animals | Test Article | Dose Volume per Hemisphere (uL) | Necropsy Study Day |
|---|---|---|---|---|---|
| 1 | Substantia Nigra | 3 | AAV | 25 - 50 | 29 +/- 4 |
| 2 | Substantia Nigra + Putamen | 4 | AAV | 25-50; 150 - 250 | 29 +/- 4 |
| 3 | Substantia Nigra + Putamen | 2 | Control | 25 - 50; 150 - 250 | 29 +/- 4 |
| 4 | Putamen | 3 | AAV | 150 - 200 | 29 +/- 4 |
Cost / Schedule / Proposal Fragments
The Northern Biomedical Research (NBR) costs we estimated at $750k.
We would add device related costs which will be in the range $125k - $200k depending on final study design.
~550,000 USD (~300K in FY2022 + ~250K in FY2023)
for in-live portion
Prescreen -> D1: Dosing -> D29: necropsy
We propose neurosurgery with AAV9 and/or AAV5, testing direct delivery to SN and putamen, and combination of both. InterimThe last line is cut off at the bottom of the photo.
Uncertain Spans
- Top cohort / ClinGen content is a continuation from
20240722_182033; use the previous capture plus this one as combined evidence before de-duplicating. - Sample-size heatmap cell values are small and remain image-primary; axes, per-arm interpretation, repeated-measures assumption, and detected reduction values should be checked before structured numeric extraction.