MC1 PET / PRKN Natural History Study Proposal And Sample Size Estimation
MJFF / PPMI / Leuven MC1 PET Proposal Notes
Eberling / Timeline Fragment
Eberling
- Timeline: 2022 or 2023
- Next step: meeting Ken (Kenneth Marek)
- f/u chat with Jaya: let's add neuromelanin-MRIDated Notes
| Date | Person / label | Stable text |
|---|---|---|
20220907 | Cristian | Following up with Jamie Eberling, the note says there may be more flexibility than expected. It may only need the consortium steering committee endorsement to get support, and not every company in the consortium would need to agree. PPMI had interest in exploring MC1 in 2023, and a PRKN sub-study could be included there. |
20221030 | unlabeled | A proposal to the MJFF consortium meeting Dec. 5th is mentioned. The note says confidentiality should be sorted before leadership and that previous in vitro and animal-model evidence indicates PRKN dysfunction affects mitochondrial function and quantity. |
20221030 | unlabeled | Action fragments: speak confidentially to Jamie and Ken; CRO sponsored study with Invicro considered; subject accessibility is the key element. |
20230110 | Cristian | Prepare a plan / slide deck for Jamie and Ken; assess interest; determine what resources MJFF is willing to contribute; seek MJFF support to identify subjects with PRKN; present to the MJFF steering committee. |
20230110 | Cristian | Koen Van Laere at Leuven University in Belgium is mentioned as a possible collaborator for a PRKN study; they have a Parkinson’s clinic. |
20230117 | unlabeled | MJFF might fund a large portion of the study through the industry consortium; they might help transfer MC1 tracer synthesis to Leuven from London; prepare a study plan with required funding. |
20230117 | [slide prep] | Therapeutic hypothesis: halt/reversal -> Longi PD/HC; Cost sharing?; Patient recruitment (below KOLs). |
20230120 | unlabeled | Rationale, DA already, NDU MEETING, ndu MEETING presentation, pt recruitment PPMI (handful). Preclinical study note mentions confusing fibroblast results, PM 8-10 w, and Sarah Sheik (budget). |
PRKN Longitudinal Observational Study Proposal
The study-design table is partly cut off at the bottom.
| Field | Text |
|---|---|
Objectives | Identify biomarkers that track clinical progression in individuals with symptomatic early-stage Parkinsonism due to mutations in PRKN. Develop a "trial ready cohort" of PRKN-PD patients for future clinical trials. |
Study Design | Prospective longitudinal observational study; Part 1 compares HC to PRKN-PD; Part 2 is longitudinal. |
Subject Population and Sample Size | N ~ 25 individuals with 10 healthy control and 15 symptomatic PD within 2 years of symptom onset. Biallelic PRKN-PD mutation carriers, confirmed LOF mutation in PRKN. Ideally presymptomatic subjects would be included, but identification may be challenging. |
Period of Evaluation | 3 - 5 years, with yearly assessments. |
Endpoints | UPDRS; Unified dyskinesia rating scale; Global Dystonia Rating Scale; in-clinic digital motor assessments of tremor, gait, bradykinesia, with dystonia and speech also considered. |
Endpoints | 7T MRI with volumetric, neuromelanin, and DTI evaluation of the substantia nigra. |
Endpoints | DaT SPECT or VMAT2 PET; 31P MRS of substantia nigra if feasible; Parkin PET if available; MC1 PET and FDG-PET under consideration. |
Endpoints | Plasma biomarkers including NfL, dopamine metabolites, lactate, pyruvate, creatine-kinase, and sphingomyelins. |
Endpoints | CSF biomarkers including pS65-Ub, VDAC, phosphorylated Parkin, TOM40/20, mtDNA, cytochrome C, and IL-6 levels. |
KOL Notes And Cohort Feasibility
Roy Alcalay
Fragment:
[20210812]
In their trial [venglustat?], Sanofi allowed people who were already treated,
at different stages and with variants that don't cause Gaucher.
Ideally, if you have enough people, you'd limit the heterogeneity,
limit it to people with no cognitive impairment or mild cognitive changes,
and not include variants that don't cause Gaucher.Christine Klein
Fragments:
GP2?
[meeting 20220708 with imaging expert (Norbert Brueggemann).] psp as negative control.
Peripheral BM: 13C-glucose ... CK in DBS by mass spec -> increased in parkin-pd (?),
but variability 많아 not a good BM, 20230519 mjf
mt workshop 에 발표. CK 아니라 gluconeogenesis 라는 marker 제시 (publication 했다 함)
MIGB scan도 차이없더라.
Animal model: vitamin k2 in Fly model (pink1 mutation?) -> ↓ motor parkinsonian,
this triggered our clinical trial (with vit k2?)
Inflammation: C Klein 2018 nature: ↓ parkin -> ↑ STING -> ↑ il-6Cohorts
Leading note:
is feasible. <2 is not feasible. <5 may be feasible| Cohort / source | Fragments |
|---|---|
PDGene | US; Roy Alcalay; 20230327 PD GENE_Takeda; PRKN Homozygotes/compound heterozygotes: n=45; PRKN heterozygous: n=79. |
GP2 | {Vollstedt, 2023 #2260}; (www.gp2.org); Europe; Christine Klein; Less active than PDGene for patient recruitment. |
Ropad/Centogene | EUROPE; ROPAD NCT03866603; Europe & Israel; funded by Denali, Centogene; focus is LRRK2, but everyone is genotyped to PRKN as well; (gabi, 20220927) >200. |
Koen Van Laere, Wim Vandenberghe | Leuven University in Belgium; identified 5 subjects with PRKN mutations; willing to talk to explore running a study; talk to Jamie to see if this can integrate with MJFF. |
MJF? | Korean note: MJF(에 PRKN 추가하라고 부탁해라). |
PPMI | 6 (?) by Jaya. |
Arndt Rolfs (Arcensus) | (gabi, 20220927): another person for finding patients could be Arndt Rolfs; he has a long history with Gaucher. |
ClinGen / PDGene / Centogene / Roy Alcalay Documents
ClinGen / PRKN Curation Note
We started curating Parkinson's genes through ClinGen, which is an NIH supported platform that is recognized by the FDA.
a major effort needs to be invested in calling PRKN mutations versus VUS.
However, LRRK2 and GBA were prioritized.
It would be extremely important if a group of scientists would curate the variants to determine which is a VUS and which is a mutation.
It may significantly change the number of carriers.
Please take a look at the panel we assembled:
https://clinicalgenome.org/affiliation/40079/.
There are researchers (Ed Fon) who are on the panel exactly for PRKN.URL:
http://www.pdgene.org/view?poly=rs3734464Side notes:
MDS gene
Denali has announced a strategic collaboration with a gene diagnostic lab, Centogene, 불량,Centogene / ROPAD fragment:
CENTOGENE's Biodatabank contains N~140 genetically confirmed PRKN-PD patients
that were enrolled into the ROPAD study.Roy Alcalay Document Links
Lead-in:
I started to put together some questions for our Monday meeting with Roy Alcalay:Linked filenames:
Dr. Roy Alcalay discussion 2-20-23.docx
conversation with Dr Alcalay 8-12-2021.docx
Questions for Roy Alcalay 1-4-2022 meeting.docx
Meeting with Roy Alcalay June 28 2022.docxSample Size Estimation
The sample-size section is partially cut off at the bottom of the photo.
Heading and question fragments:
Sample size estimation
- What does the power or magnitude of effect need to be in order to see an effect in a sample size of 15-25 people?
- Using VMAT2 (a sensitive imaging biomarker), the change of VMAT2 signal over 2 YEARS is as follows:
Mean: -19.6%, SD 9.5%, % COV: 48% (=9.5/19.6)
- What sample size is needed to see a change over 1 year? Over 2 years?
To check the below sample size is per arm,? repeated-measures t-test90% Power / One-Sided p-value < 0.05
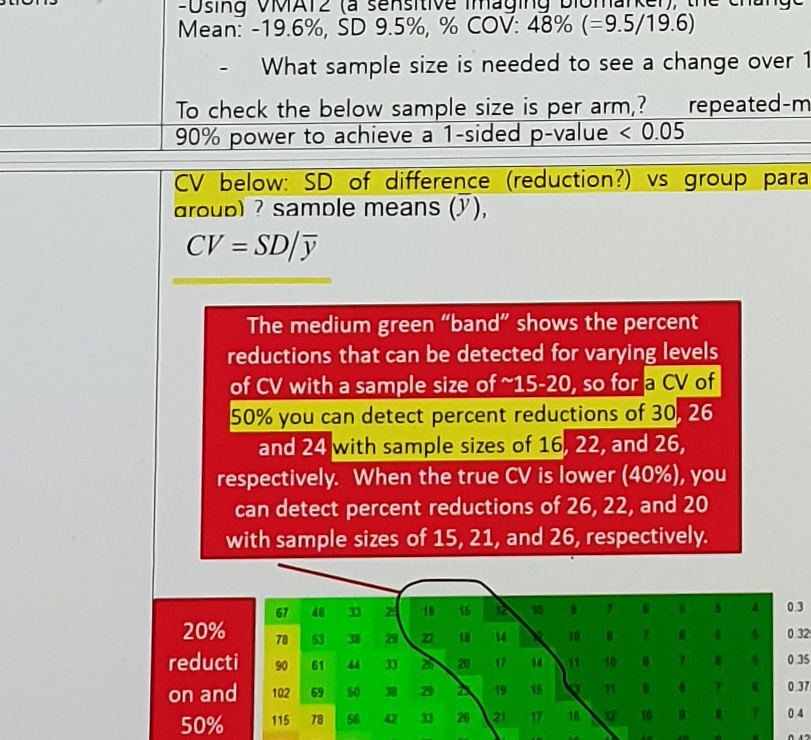

Formula / assumption text:
90% power to achieve a 1-sided p-value < 0.05
CV below: SD of difference (reduction?) vs group parameter (each group?) sample means (y),
CV = SD/yRed-box note:
The medium green "band" shows the percent reductions that can be detected for varying levels
of CV with a sample size of ~15-20, so for a CV of 50% you can detect percent reductions of 30, 26
and 24 with sample sizes of 16, 22, and 26, respectively.
When the true CV is lower (40%), you can detect percent reductions of 26, 22, and 20
with sample sizes of 15, 21, and 26, respectively.80% Power / One-Sided Alpha = 0.10
Assumption:
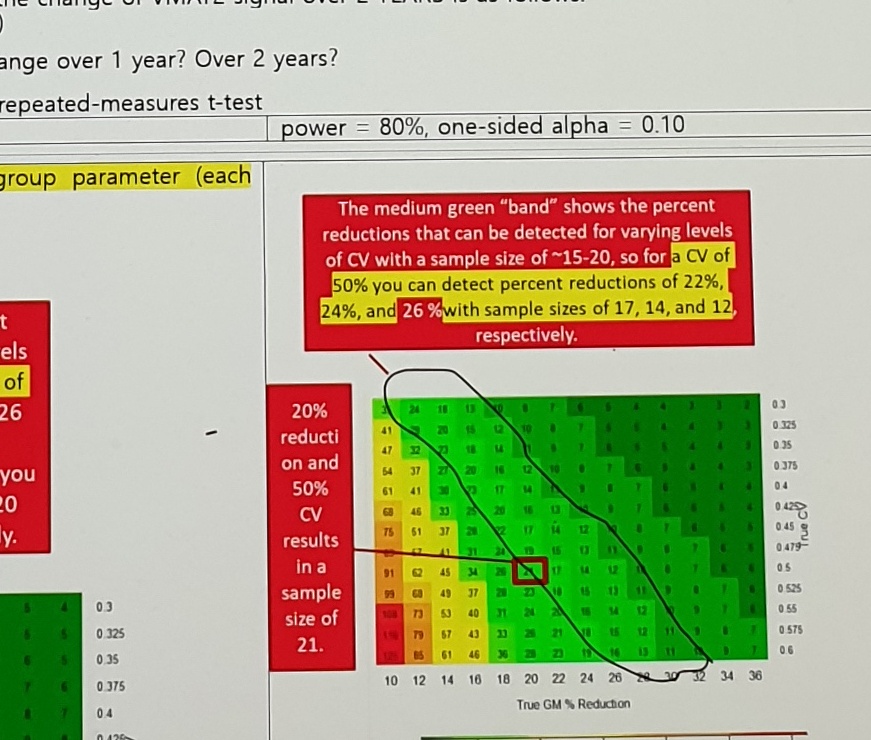
power = 80%, one-sided alpha = 0.10Red-box note:
The medium green "band" shows the percent reductions that can be detected for varying levels
of CV with a sample size of ~15-20, so for a CV of 50% you can detect percent reductions of 22%,
24%, and 26% with sample sizes of 17, 14, and 12, respectively.Left annotation:
20% reduction and 50% CV results in a sample size of 21.Uncertain Spans
- The natural-history/cohort material spans adjacent scroll positions; use neighboring photos before treating this page as a complete section.
- Sample-size heatmap cell values are image-primary; use the embedded figures before extracting exact per-cell numeric values.