cGAS: Senescence
cGAS Evidence Table Continuation
Paraquat / iPSC Correction Rows
| row / model | evidence and pathway notes |
|---|---|
Mice: Paraquat (inhouse) | ↑ cGAS (trend) (midbrain); ↑ cGAS; = cGAMP (under CCCP); = STING; = TBK1; = IRF3; = IFNβ1, ISG15; ↑ cleaved caspase-3. |
| right-side note | {Sliter, 2018 #699} ↑ serum IL-6, IL-1b, CCL2, CCL4 in PINK1 hetero, PARK2 heteroz, PARK2 homoz, sPD (there is no PINK1 homo) (IFNβ1는 비인급?) implication: cytokine alteration is not sufficient to cause disease. Not a DR marker? |
Correction / iPS: Parkin KO iPSC-DAn | (cf: basal level은 ↓); STING not detected; No effect of cGASi (tool) on IFNβ1 or ISG15; No effect of cGASi (tool) on CC3. |
Sliter 2018 Mouse / STING Deletion Table
| marker / readout | Following EE | PARKIN-/- | Pink1-/- | Prkn-/-; STING... | Pink1-/-; STING... |
|---|---|---|---|---|---|
| table context | {Sliter, 2018 #699} acute exhaustive exercise EE-induced mitochondrial stress model | Sting deletion model | Sting deletion model | ||
pS65-ub | Yes | No increase | |||
Mt-keima | yes | No increase | |||
Serum cytokines | Yes | ↑: IL-6, IFNβ1, IL-12(p70), IL-13, CXCL1, CCL2 or CCL4, ↑ fever (Fig. 1d, e, Extended Data Fig. 1e-i). | No increase in the cytokines, (Fig. 2c, d; Extended Data Fig. 3d-i), no fever | ||
Serum cytokines | Yes | ↑ IL-1b Fig 1e, extended data fig. 1j | |||
serum mito DNA | Yes | ↑ (fig3, by PCR) | NA | ↑ | = |
cGAMP (Heart, LCMS) | yes | ↑ | ↑ (extended data fig 4a) |
{Sliter, 2018 #699} STING deletion -> ↓ Inflammation
(resulting from either exhaustive exercise or mtDNA mutation) (in PARKIN-Deficient mice?)
{Sliter, 2018 #699} STING deletion also rescued the loss of dopaminergic neurons and motor defects observed in Parkin-deficient mutator mice, which accumulate damaged mitochondrial DNA,
Cf) 2015 Gao: Even deletion of just one allele of cGas largely rescued the Trex1-/- phenotypes of miceBiomarker / Readout Notes
| readout / question | note |
|---|---|
| dsDNA | can we select high dsDNA segmentation in PD patients? |
cGAS protein | 20200930 Masato: Its expression level is low normal brain, so It should take time to setup LCMS. |
cGAMP | |
| ISG | ISG는 너무 다양하니 부적합하겠군 |
| cGAS PET | Cgas PET: Activation level might be different in the same patient throughout the day. cGAS is not activated in HV |
| cytokine candidates | Type I interferon?; TNF-a and IL-12 |
cGAS-STING Pathway Figure
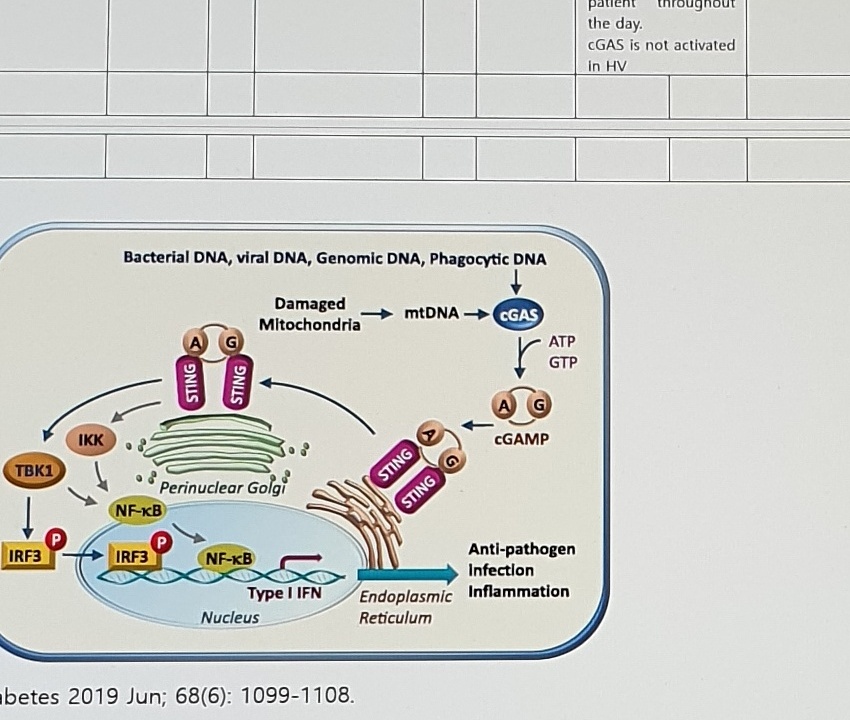
Bacterial DNA, viral DNA, Genomic DNA, Phagocytic DNA
Damaged Mitochondria -> mtDNA -> cGAS
ATP / GTP -> cGAMP
STING
Perinuclear Golgi
TBK1
IKK
IRF3
NF-kB
Type I IFN
Nucleus
Endoplasmic Reticulum
Anti-pathogen Infection InflammationDiabetes 2019 Jun; 68(6): 1099-1108.Mito Damage + Senescence Table
[mito damage + senescence]| trigger | Exp model | mitochondrial damage | mitochondrial DNA release | cGAS-STING | senescence / type I IFN | inflammation | neuronal loss |
|---|---|---|---|---|---|---|---|
Parkin loss | PARK2 KO iPSC-DAn | Mitochondrial damage | Mitochondrial DNA release | Activation of cGAS-STING pathway | ↑ type I interferon, | Neuroinflammation | Neuronal loss |
Stress (eg paraquat) | paraquat-injected mouse brain | Mitochondrial damage -> ↑ ROS; (cf: there have bee[n] (Fukushima, 2002 #1018) no reports on mitochondrial DNA damage by paraquat) | Nuclear dsDNA release | Activation of cGAS-STING pathway | ↑ senescence of astrocyte (↑ p16INK4a); SASP (MMP-3 and the pro-inflammatory cytokines interleukin-6 (IL-6), IL-1α, and IL-8) | Neuroinflammation |
Executive Summary cGAS
Executive summary cGAS
https://mytakeda.sharepoint.com/sites/cGASOur compound binds activated Cgas, non-activated cGAS에도 붙는지는 확인중- cGAS protein expression:
cGAS was relatively highly expressed in human heart, lung, kidney, skin, and skeletal muscle (but low in brain) (Niigata univ collaboration 20210129)
Senescence
Basic Hypothesis
Senescent cells damage neighboring cells by releasing cytokines, chemokines, proteases and growth factors (senescence-associated secretory phenotype (SASP))Evidence in PD
(Chinta, 2018 #936).Hypothesis:
Stress (eg paraquat) -> injured/dying cells release dsDNA (self DNA) into the cytoplasm
(js: I can't find this in Chinta 2018) -> recognized by Cgas in phagocytic monocytes and/or macrophages
-> ↑ STING (stimulator of interferon genes) in ER in macrophage
-> ↑ senescence of astrocyte (ie not dividing)
-> ↑ pro-inflammatory factors
-> ↑ neurodegenerationEvidence:
postmortem brain samples of sPD patients.
SNpc:
↑ senescence marker p16INK4a(senescent cells, both in vivo and in vitro, often express p16(Ink4a))↑ several SASP factors included the protease MMP-3 and the pro-inflammatory cytokines interleukin-6 (IL-6), IL-1α, and IL-8
Astrocytes in SNpc:
↓ lamin B1(A reduced nuclear level of lamin B1, detectable by immunostaining in intact tissue, is an established senescence-associated marker)- Experiment:
- In vitro
In Vitro
Paraquat:
Paraquat -> (human astrocytes derived from induced pluripotent stem cells (iPSCs))↓ Proliferation (↓ bromo-deoxyuridine (BrdU)-labeled cells)hallmarks of senescence, including increased senescence-associated β-galactosidase (SA-β-gal) activity (Dimri et al., 1995) (Figure 2B), increased levels of p16INK4a (Beausé jour et al., 2003) mRNA and protein (Figures 2C and 2D), and increased numbers of 53BP1 foci, the latter indicative of DNA-damage signaling, which is a potent stimulator of the SASP (Beausé jour et al., 2003)the astrocytes to secrete certain proteins--including proinflammatory factors IL-6
Conditioned medium:
CM from PQ-induced senescent astrocytes-> ↓ DAergic neuronal viability
IL-6:
Just a trend toward ↓ DAergic neuronal viability
Mouse Model
the scientists made use of a mouse model that allows for the selective deletion of senescent cells, and injected them with paraquat over the course of a few weeks. Similar to human Parkinson's disease, this resulted in the loss of dopaminergic neurons and of motor function. Most interestingly, the researchers were able to halt the progression of these symptoms by clearing all senescent cells from the substantia nigra.MOA Integrated
MOA Integrated
Normal
(mito rupture까지 가기전에) Mitophagy로 mitoch cleared
No Mito rupture
Mitochon
↑ ROS release
Cell
Aging, exercise,, stress, mito
Parkin
mitophagy장애Uncertain Spans
| location | text/status | reason |
|---|---|---|
| nav path | cGAS > Senescence | The navigation pane visibly highlights Senescence; the status bar still shows generic Supplement / Positive vs Negative entries, so nav/body evidence is stronger. |
| page label | Page 44 of 50 | Apple Vision, PaddleOCR, and visible status bar agree, but nearby synthesized pages have shown document/page-count inconsistencies. |
| Paraquat row | = STING, = TBK1, = IRF3, ↑ cleaved caspase-3 | Narrow table cells and line wrapping make exact equality/arrow relation high-risk. |
| right-side Sliter note | (IFNβ1는 비인급?) | Korean note is red and partly blurred. |
| STING deletion genotype labels | Prkn-/-; STING..., Pink1-/-; STING... | Superscript/subscript genotype notation is too small to fully resolve. |
| Sliter table symbols | PARKIN-/- cells for pS65-ub and Mt-keima | The cells show small symbols, but not enough to confidently transcribe as +, -, or another mark. |
| dsDNA question | high dsDNA segmentation in PD patients? | The word segmentation is split over several lines and could require domain review. |
| cGAS protein note | low normal brain | Visible wording may mean low in normal brain; left as typed/visible. |
| mito damage table | Fukushima, 2002 #1018; bee[n] no reports | The line is cramped; bracketed n marks an inferred missing character from line wrap. |
| senescence paragraph | Beausé jour / Beauséjour; SA-β-gal; 53BP1 | Citation spelling and Greek letters are high-risk OCR items. |
| MOA Integrated | bottom table | Only the first rows are visible. |